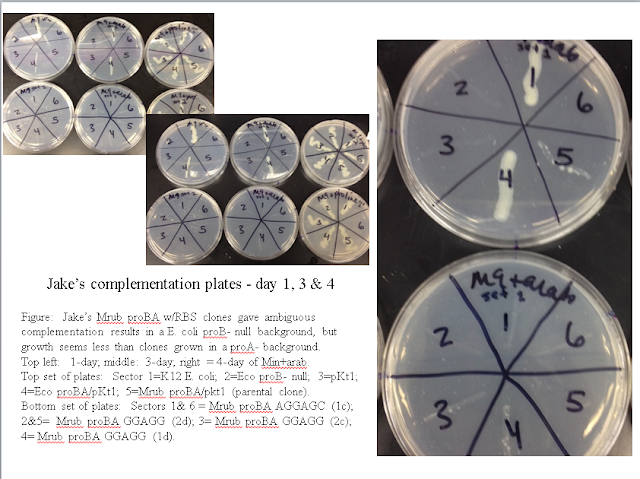
In an E. coli proA null background, both the positive controls (E. coli proBA and M. ruber proBA) and the experimental (M. ruber proBA with new RBS) complemented (see figure below). Growth was slow for both M. ruber clones, however. This is contrary to our observations from the winter term Molecular Genetics course, which saw no growth for the non-mutated M. ruber proBA clone. It's clear, however, that the new RBS did not give the mutated clone an advantage under these conditions.
In an E. coli proB null background, very little complementation of either M. ruber clone was observed (see image below).
We successfully inserted two different RBS sequences (as confirmed by sequencing), but maybe we don't have the right sequence. The next step is to try a third RBS sequence. I'm still thinking about why we see different complementation results for the E. coli proB vs. proA null strain. As described in earlier videos, proB and proA form a multi-protein complex that is required for proB function but not required for proA function.